Out of 180,108 assembled transcripts, 13,636 were specific to the Svevo cultivar as compared to the only other reference transcriptome available for durum, thus contributing to the identification of the tetraploid wheat pan-transcriptome. Our study updates and expands the de novo transcriptome reference assembly available for durum wheat. The transcriptomes of 13 durum wheat cultivars spanning the breeding period from 1969 to 2005 were analysed for SNP diversity, leading to 95,358 non-rare, hemi-SNPs shared among two or more cultivars and 33,747 locus-specific (diploid inheritance) SNPs. We observed greater expression divergence between A and B homoeologs in grains rather than in leaves and roots. 
urartu transcripts and inferring homoeolog-specific sequences. The presence of differential expression between the A- and B-homoeolog copies of the durum wheat tetraploid genome was ascertained by phase reconstruction of polymorphic sites based on the T.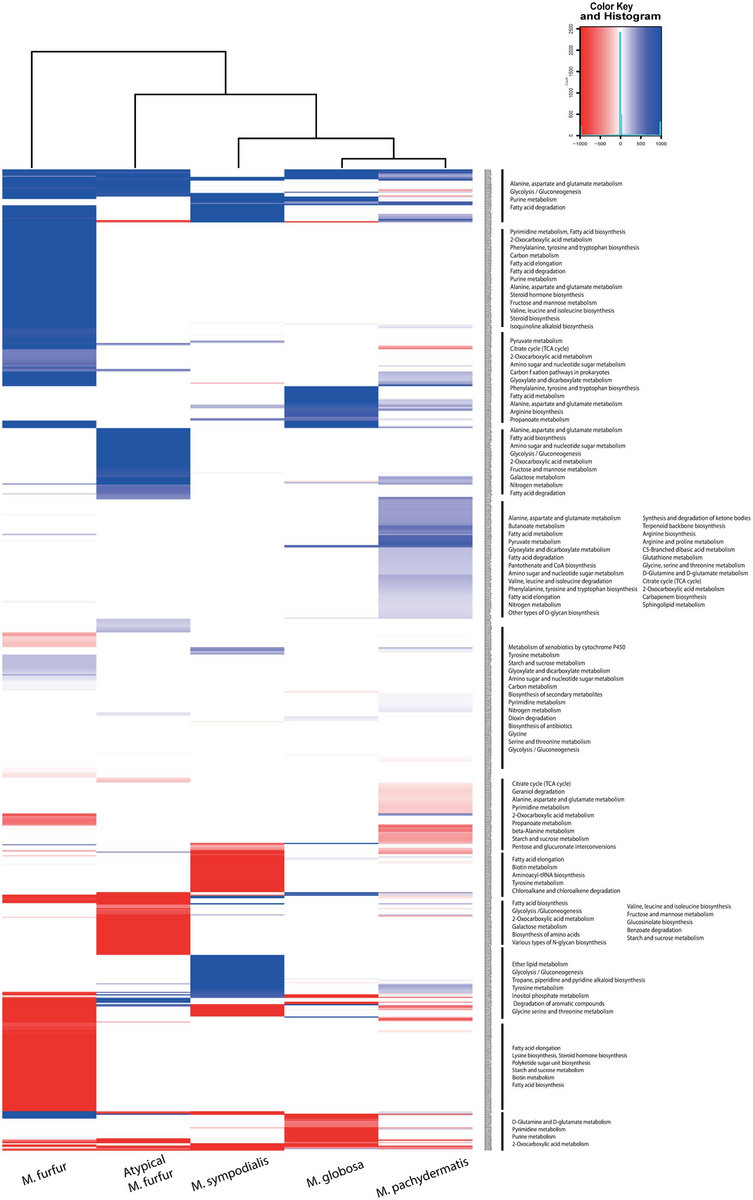 
The functional annotation was completed by means of a combination of complementary software. Alignment against the transcriptome of nine plant species identified 43% of transcripts with homology to at least one reference transcriptome. The transcriptome of four tissues/organs (shoots and roots at the seedling stage, reproductive organs and developing grains) was assembled de novo, yielding 180,108 contigs, with a N50 length of 1121 bp and mean contig length of 883 bp. We report on the de novo transcriptome assembly of durum wheat cultivar ‘Svevo’. A deeper characterisation of the molecular and functional diversity of the durum wheat transcriptome will be instrumental to more effectively harness its genetic diversity. Future breeding efforts to enhance yield potential and climate resilience will increasingly rely on genomics-based approaches to identify and select beneficial alleles. Husnot) is an important crop which provides the raw material for pasta production and a valuable source of genetic diversity for breeding hexaploid wheat ( Triticum aestivum L.). Our aim is to provide a quick start guide to the nonexpert researchers for NGS-based transcriptome analysis.The tetraploid durum wheat ( Triticum turgidum L. Further, we describe a method for using RNA-seq to characterize the transcriptome of a plant species, taking the example of a legume crop plant chickpea. Here, first we outline various important issues from experimental design to data analysis, including various strategies of transcriptome assembly, which need substantial consideration for a successful RNA-seq experiment. Further, the assembly of millions and billions of RNA-seq reads to construct the complete transcriptome poses great informatics challenges. Although becoming cheaper, transcriptome sequencing still remains an expensive endeavor. The transcriptome sequencing of an organism provides quick insights into the gene space, opportunity to isolate genes of interest, development of functional markers, quantitation of gene expression, and comparative genomic studies. Sequencing of mRNA using next-generation sequencing (NGS) technologies (RNA-seq) has the potential to reveal unprecedented complexity of the transcriptomes. The integration of data from genomics and proteomics analysis allows for the composition of interactomes, elucidating systems wide impacts resulting from disruption of the CCM signaling complex (CSC). Here we describe the methods currently being used to evaluate CCM-deficient strains in human brain microvascular endothelial cells (HBMVEC), zebrafish embryos as well as in vivo mouse model to evaluate impacts on various signaling cascades resulting from deficiencies in KRIT1 (CCM1), MGC4607 (CCM2), and PDCD10 (CCM3). Proteomics facilitates an understanding of mechanisms being altered at the translational level allowing for an understanding of multiple layers of regulation occurring, elucidating discrepancies between what is seen at the RNA level compared to what is translated to a functional protein. Genomics, performed through RNA-seq, allows the user to evaluate alterations at the transcription level, oftentimes more sensitive than other types of analysis, especially when attempting to understand lack of observation of an expected phenotype. Omics research has garnered popularity recently to integrate in-depth analysis of alterations at the molecular level to elucidate observable phenotypes resulting from knockdown/knockout models.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
